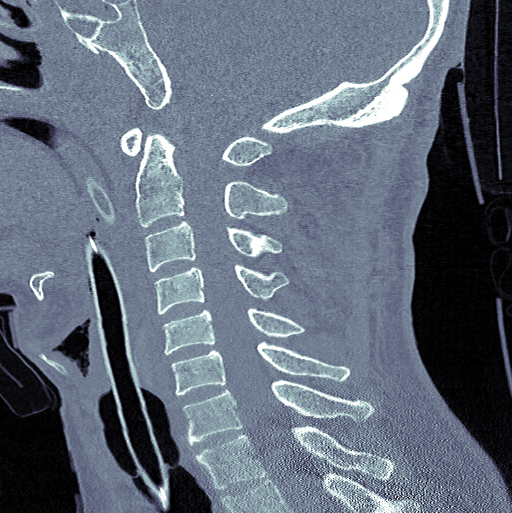
Cervical Spine Fractures
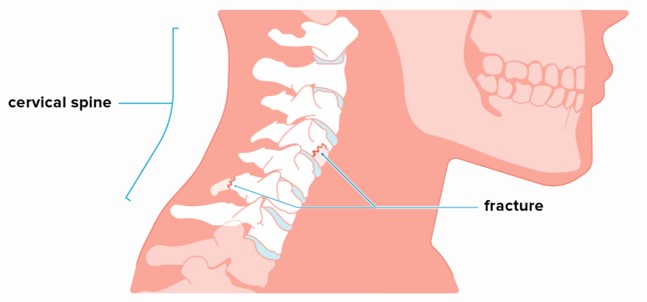
Annually in the USA, an estimated 1.5 million vertebral compression fractures occur — the most common located in the cervical spine where only 25–33% of incident radiographically identified vertebral fractures are clinically diagnosed. Quickly detecting and determining the location of vertebral fractures is essential to prevent neurologic deterioration and paralysis after trauma.
Objective:
- Train deep learning algorithms based on 2019 patients CT images
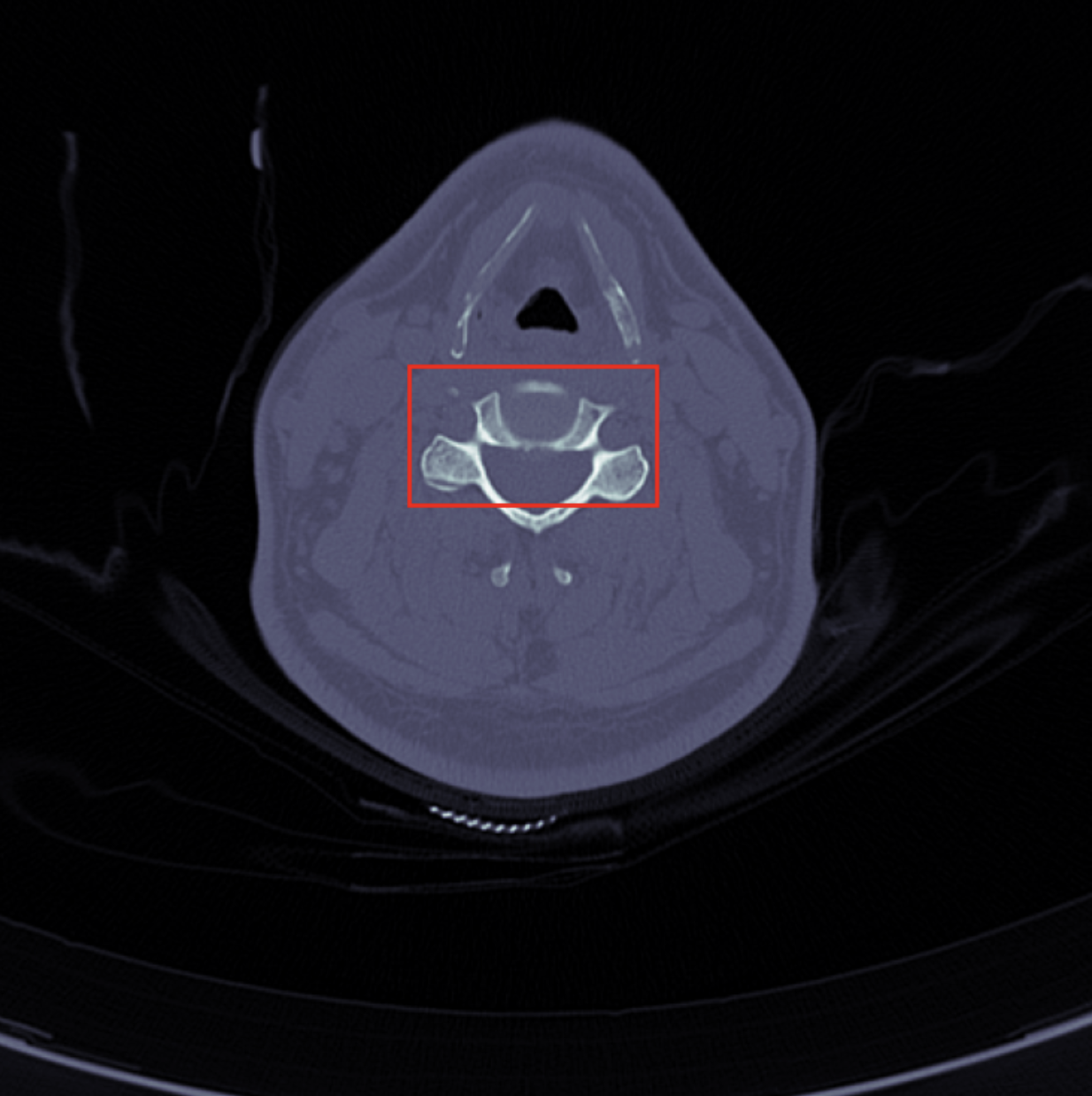
- Detect possible cervical fracture based on CT axial images
- Locate the fratures by bounding boxes
- Evaluate deep learning model performance
Our Mission
Quicker, better, more accurate diagnosis to save lives.
Minimal Viable Product
Develop deep learning algorithms to detect if there are cervical fractures from CT images of the cervical spine.
Main Features
Detect cervical spine fracture and locate the fractures.
Product Vision
Help every radiologist quickly identify the cervical spine fractures within a few minutes.
Market Size
The global spinal imaging market size was valued at $1.6 billion in 2019 with expect growth rate of 5.2% from 2020 to 2027.
Customers
Potential customers are hospitals, diagnostic imaging centers, and ambulatory care centers.
Competitors
The competitors are other AI models. So far, only one company luanched their commercial model in and the market is not saturated at all.
Demo

 NeckFrac with Sagittal Views NeckFrac with Bounding Boxes
NeckFrac with Sagittal Views NeckFrac with Bounding Boxes
Dataset

Public data provided by RSNA (Radiological Society of North America)
- Collected imaging data sourced from 12 sites on six continents.
- Includes over 2,000 CT studies with diagnosed cervical spine fractures and an approximately equal number of negative studies.
- Data size (343 GB), each patient has around 300 images for CT scans.
- Experts labeled the annotation to indicate the presence, vertebral level and location of any cervical spine fractures.
Pipeline

Model Architecture
Model I: EfficientNet

-
It’s all about Scale! Convolutional neural networks need to be scaled up to achieve better model performance; however, scaling is quite random among the conventional techniques of model scaling. Some models scale depth-wise, some scales widthwise and some takes in images of a larger resolution to get better results. These techniques require manual tuning and many person-hours with in little or no improvement in performance. EfficientNet uses a technique called compound coefficient to scale up models in a simple but effective manner. Instead of randomly scaling up width, depth or resolution, the compound scaling method is based on the idea of balancing dimensions of width, depth, and resolution by scaling with a constant ratio. The intuition for EfficientNet is that the network needs more layers to increase the receptive field and more channels to capture more fine-grained patterns on the bigger image.

EfficientNet Cervical Spine Fracture Detection Model Implementation
- Pre-processing:
- Hounsfield Units rescale to sharpen the intensity of bone, fat tissue, etc.
- Normalization with max bound and min bound. Max bound = 2050 and min bound = 150 to focus on portion of the scan that is bone tissue.
- Rescale image to improve the training speed. We resized the image from 512x512 pixel to 128x128 pixel.
- Calculate class weight:
- To treat the imbalanced dataset problem, we applied calculated class weight to this model, so the underrepresented class has a much higher error than the majority class.
- 8 EfficientNet Models:
- We experimented with different class weight to the EfficientNet models: due to different class weight, training each binary classification separately with adjusted class weight to each vertebrate label yield the best result.
- We applied 2-fold cross validation and trained by batch with batch size of 150 patient and learning rate of 0.00001. The total training time is around 9 hrs.
- Inference:
- We inferred 500 patients with the EfficientNet Model with average inference time of 58 secs on inferring 8 models all at once.
Model II: 2.5D UNet + biGRU
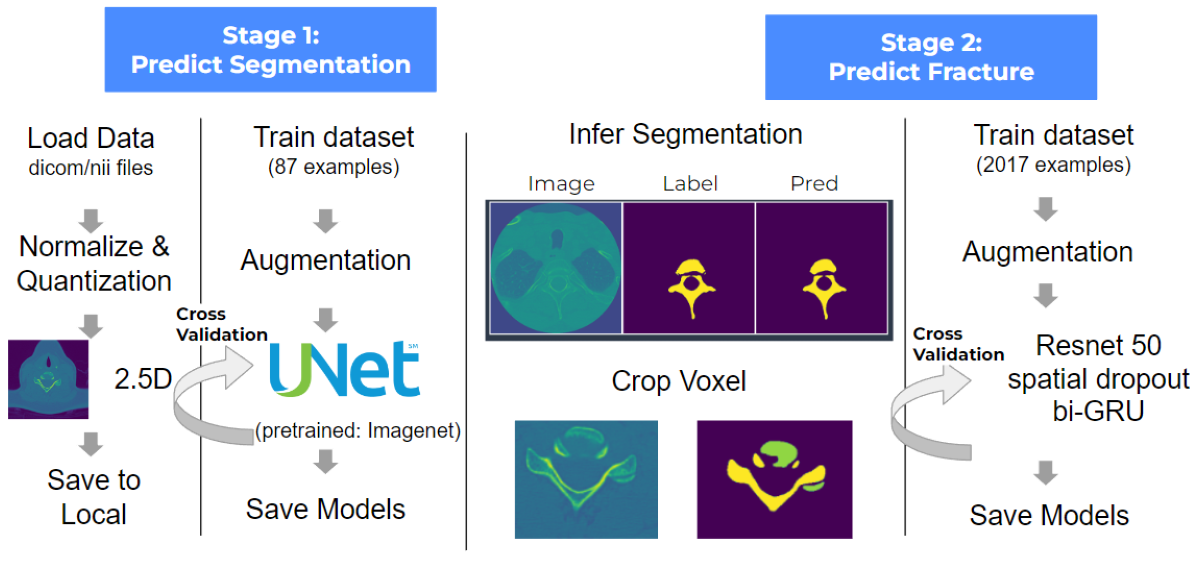
- Prepare 2.5D slices.
- Train a model to predict segmentation.
- Use the best model from step 2 to infer segmentation for all rest of train images and only crop out the vertebrae.
- Train the model to predict fractures on each vertebra.
Segmentation is the image pixel labeled as C1, C2, etc. In the dataset, only 87 patients have labeled segmentation, which were used to train during stage 1. Before training, we normalized the pixel array and changed the type as integer 8 to speed the computation. Then we preprocessed the image into 2.5d and saved them locally. Here 2.5D means that each 2D slice in a vertebrae sample has the information of the adjacent slices.
Then, the 2.5D slices were augmented with random flip, rotation, bur, and were passed into UNet with image and labels. We did 5-fold cross validation and implemented the early stop, and after 9 epochs, the best model was returned.
Stage 2:We used the best model from stage 1 to infer segmentation for the rest of the train set. Crop the images and focus on vertebrae only. After cropping vertebrae, we did a similar augmentation as step 1, and passed the images into CNN with Resnet 50 backbone and bi-GRU with spatial dropout. We did 5-fold cross validation and implemented the early stop, and after 11 epochs, we got our best model. For inference, this model takes 40s for each patient to get the fracture result.
Performance
Ensemble Model Performance


Error Analysis

Business Impact

Compared to Small et al. (2021), we outperform radiologists and an industry competitor (AIDOC) on recall/sensitivity and inference time. Our model would be best utilized in situations where the radiologist would like to sift through predicted negatives quickly and focus on predicted positives.
Future Work
- Labeling stretched images for better error analysis
- Incorporating soft tissue into model to identify fracture
- Combine three anatomical planes - axial, sagittal, and coronal - for modeling
- Designing a model API for use in CT imaging devices
- Rank the fracture severity (low, medium, high) based on probability
Portfolio
Here are the output examples from the ensemble model (click + into the images to see prediction).
Notice that better image orientation (e.g. not stretched or tilted) results in better predictions.
Our Team
Special Thanks to Dr. Puya Vahabi and Dr. Alberto Todeschini

Fengyao Luo
fengyaoluo@berkeley.edu
Weijia Li
weijia.li@berkeley.edu
Jane Hung
janehung@berkeley.edu
Minjie Xu
minjiexu@berkeley.eduAcknowledgement
We would like to express our deepest gratitude to our capstone professors: PUYA H. VAHABI and ALBERTO TODESCHINI from UC Berkeley for giving us suggestions on model training and error analysis. We would also like to acknowledge Robert Wang from AWS for providing advice for infrastructure and Joyce Shen for organizing AWS workshops. Dealing with such a large dataset has been one of the challenges that we faced, Robert was always available to help us with clarifications and answers to our questions . Last, but not least, we'd like to extend our thanks to Andrew Nguyen, DO, the radiologist in Sunrise Hospital And Medical Center - HCA Healthcare, who offered his professional knowledge on viewing patients CT scans in clinical practice and helping us understand the user scenarios in the medical environment.